Several genetic biomarkers may impact the likelihood of response or side effects of a drug. This is an issue that many clinicians frequently face.
In the Genomind PGx test report, you’ll find several pharmacodynamic (PD) genes that impact commonly utilized psychiatric medications, including genes associated with:
- Antidepressants (e.g., SLC6A4, HTR2A, MTHFR, BDNF)
- Stimulants (e.g., COMT, ADRA2A)
- Antipsychotics (e.g., DRD2, MC4R, 5HT2C)
How genes affect SSRIs, SNRIs, and other antidepressant medication efficacy
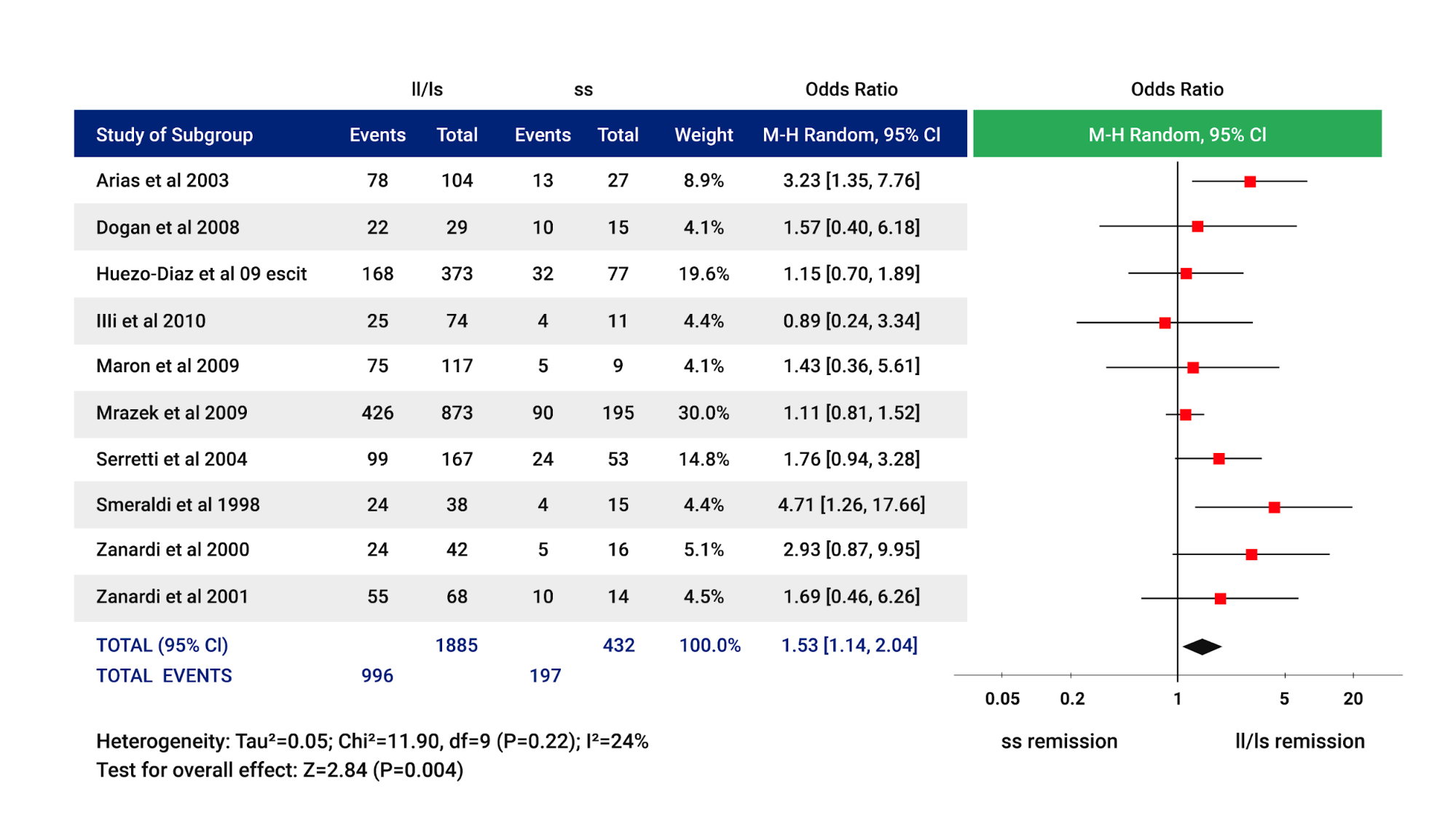
One of the most extensively studied pharmacodynamic genes in psychiatry is the serotonin transporter (SLC6A4). Caucasian patients homozygous for the short allele in SLC6A4 (i.e., S/S) are less likely to achieve remission when using SSRIs medications to treat major depressive disorder, which can be seen in the Forest plot in the meta-analysis presented in figure 1 (Porcelli et al., 2012).
This polymorphism is also associated with an increased risk for side effects in Caucasians that carry the S allele (Hu et al., 2007). Patients with the S/S genotype may be more likely to respond/remit to SNRIs and other antidepressants like bupropion. These drugs were not associated with a reduction in response/remission rates with this genotype.
Figure 1
How genes affect ADHD medication efficacy
Pediatricians and child/adolescent psychiatrists that commonly prescribed stimulants for treating symptoms of ADHD may find genes like COMT (catechol-o-methyltransferase) useful in identifying those patients more or less likely to respond to medications like amphetamine and methylphenidate.
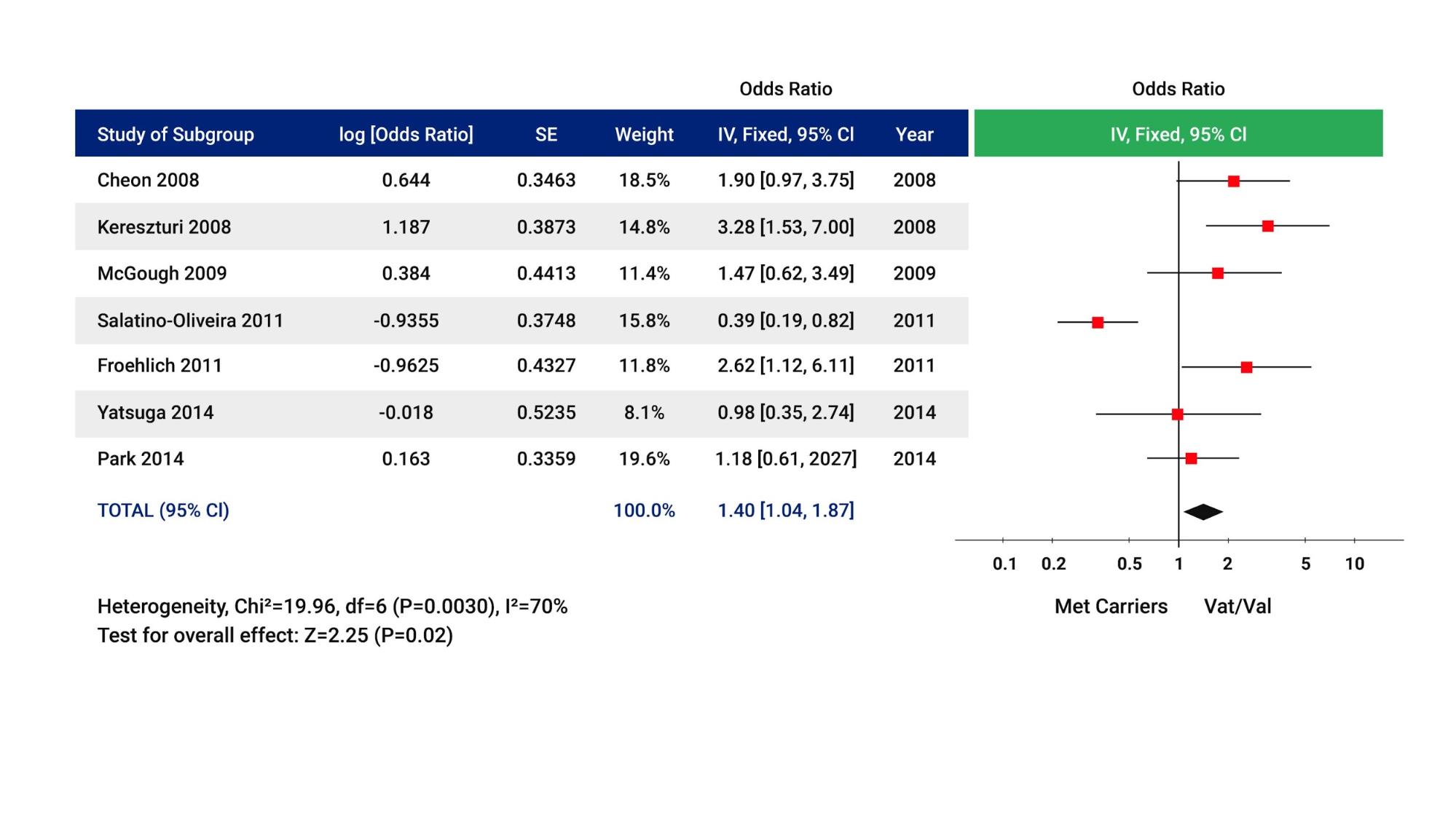
The Myer meta-analysis presented in figure 2 suggests that patients with the COMT val/val genotype may be more likely to respond to stimulants than patients carrying a ‘met’ allele. Patients carrying two copies of the met allele may not be as likely to respond to stimulant medications and may be more likely to benefit from non-stimulant alternatives (e.g., atomoxetine, clonidine, guanfacine).
Figure 2
How genes affect atypical antipsychotics treatments for schizophrenia, bipolar depression, and others
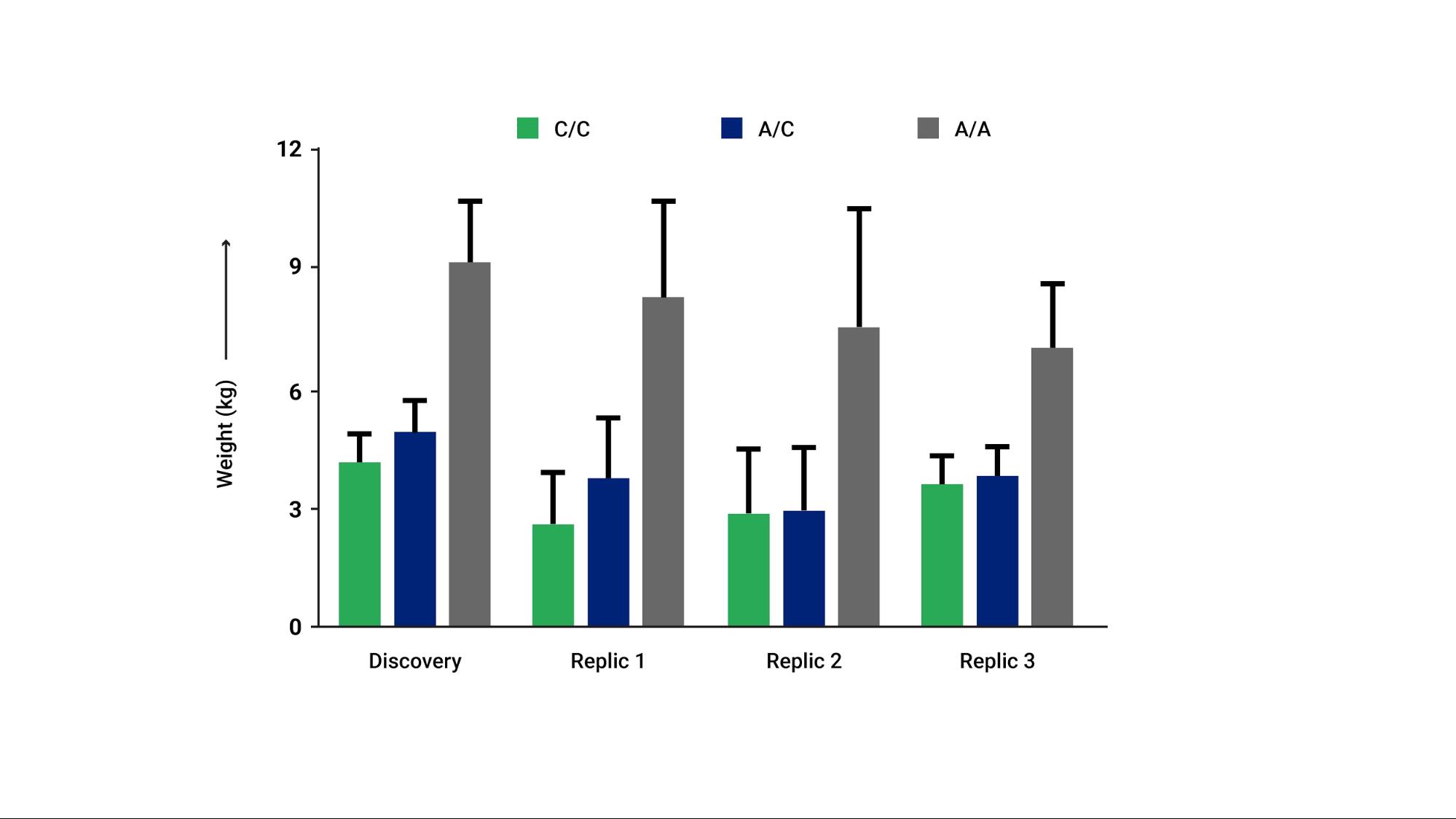
When prescribing commonly used atypical antipsychotics (e.g., aripiprazole, quetiapine, risperidone, etc.) for schizophrenia, bipolar depression, or other mood lability disorders, clinicians may want to be aware of genetic polymorphisms in the MC4R gene associated with increased weight gain. More specifically, patients with a C/C or A/C genotype gained 4.87 kg on average, while patients with the A/A genotype in MC4R gained an average of 10.03 kg. Patients with the MC4R A/A genotype may derive greater benefit from the use of more weight-neutral atypical antipsychotics.
Figure 3
In conclusion
While this section went over a few of the PD genes associated with drug response and side effect risk, the Genomind PGx test report analyzes 12 additional pharmacodynamic genes that can provide further guidance when prescribing medications for several psychiatric diagnoses, in addition to 9 pharmacokinetic genes that affect drug metabolism and exposure. These include major depressive disorder, generalized anxiety disorder, obsessive-compulsive disorder, pain, drug dependency, bipolar disorder, and schizophrenia.
Deliver targeted and personalized care with Genomind.
Register with Genomind to use our precision tools and services and help your patients get better.
